Model Validation and Performance Evaluation
Introduction
Up to now, we have trained models on a single training set and evaluated their performance on a separate test set. We used metrics such as MAE for regression and accuracy for classification to assess how well the models performed on unseen data. In earlier chapters, we also experimented with adjusting models and comparing alternatives, including hyperparameter changes, to improve performance. However, this approach is not ideal, since repeatedly using the test set for model selection effectively turns it into part of the training process. This leads to overly optimistic performance estimates and undermines its role as a final, unbiased evaluation tool.
In this chapter, we focus on validating and measuring model performance in a more principled way. We introduce resampling methods that allow us to evaluate models more reliably and systematically by repeatedly splitting the data into training and evaluation subsets. We also discuss how performance metrics should be chosen based on the goal of the analysis, since different metrics can lead to different conclusions about which model performs best.
Performance Evaluation
In the previous chapters, we highlighted that the training set should not be used to evaluate the predictive performance of a model, due to the risk of overfitting. Unless the model is very simple, such as a linear regression with only a few predictors, most models will tend to overfit the dataset on which they are trained. For this reason, we introduced the holdout method, where a subset of the data—called the test set—is kept aside to test model performance.
However, we also followed this approach mainly for simplicity. In practice, we emphasized that using the test set as a guide to tweak or improve our models makes it effectively part of the training process. This should never be the case. As Trevor Hastie, a well-known expert in machine learning, states, “ideally the test set should be kept in a vault, and be brought out only at the end of the data analysis” (Hastie et al., 2009; Lantz, 2023).
Validation Set
The simplest way to avoid using the test set during model selection is to introduce an additional subset of the original dataset. This dataset is usually called the validation set, and it is kept separate from the test set. The idea is straightforward: we train different models on the training set, evaluate their performance on the validation set, and then select the model that performs best. After identifying a satisfactory model, we evaluate it one final time on the test set to obtain an unbiased estimate of its performance.
However, even when using a validation set, the same issue can arise. A model may eventually overfit the validation set, meaning that it performs well on validation data but does not generalize well to truly unseen data. For this reason, we often rely on resampling methods, especially when the dataset is large enough to support them. Resampling methods create multiple training and testing splits from the same dataset. Instead of relying on a single split, we repeat the evaluation process several times, leading to more reliable estimates of model performance.
K-fold Cross Validation
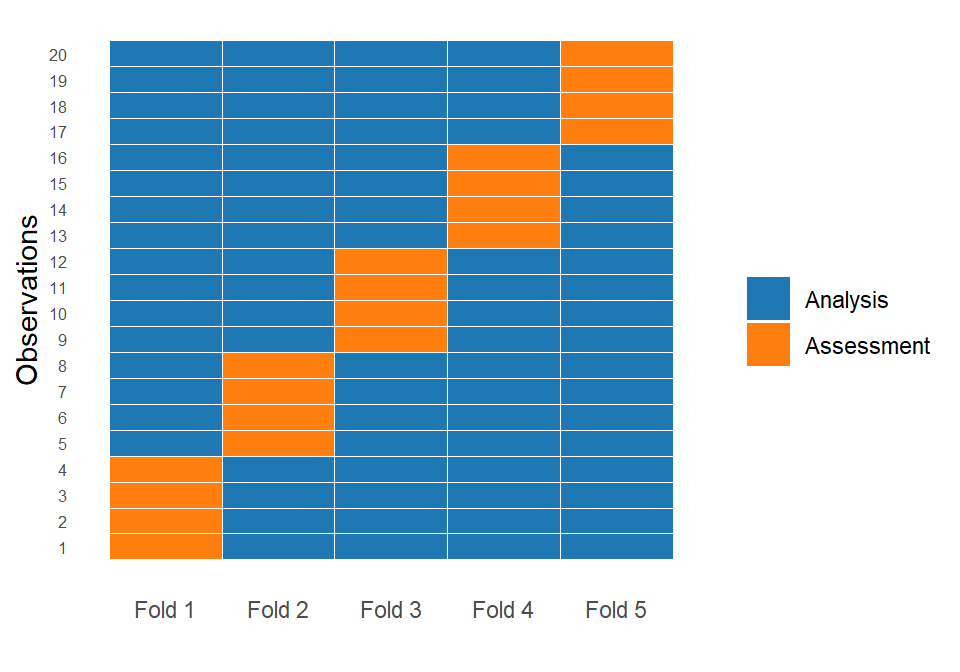
One of the most commonly used resampling techniques is k-fold cross-validation. The idea is simple: we start by splitting the training dataset into \(k\) equally sized parts, called folds. For example, if \(k = 5\), the data is divided into five folds. In each iteration, the model is trained on \(k - 1\) folds (the analysis set) and evaluated on the remaining fold (the assessment set). This process is repeated \(k\) times so that each fold serves exactly once as the assessment set.
As a result, we obtain \(k\) performance values, one from each fold. Since each value comes from a different assessment set, the result is not a single estimate but a distribution of performance values. For example, if we use MAE as the evaluation metric, we obtain \(k\) different MAE values. Instead of summarizing performance with a single number, we can describe this distribution using the mean and standard deviation, which gives us both an estimate of expected performance and information about its variability. Importantly, every observation in the training set is used exactly once for validation and multiple times in the analysis sets, ensuring that all data points contribute to both model fitting and evaluation.
In the plot above, the first fold assigns the first 4 observations to the assessment (test) set, with the remaining 16 observations used for analysis (training). In the second fold, the next group of 4 observations forms the assessment set, while all other observations are used for analysis. This procedure is repeated until each observation has been used exactly once in the assessment set across the five folds, with the remaining observations forming the corresponding training sets.
Nothing prevents us from increasing the number of folds. Common choices are 5-fold or 10-fold cross-validation. In general, a larger number of folds leads to a more detailed evaluation, but also increases computational cost.
Even though k-fold cross-validation produces multiple performance estimates that resemble a distribution, it is worth noting that these observations are not independent. The method relies on repeated reuse of the same data points across different analysis sets, which introduces dependence between the folds. As a result, the performance values obtained from each fold are correlated, since the analysis sets overlap significantly.
This means that cross-validation does not generate independent samples of model performance in a strict statistical sense. Instead, it provides a practical and highly useful approximation of model generalization performance. For this reason, cross-validation should be interpreted as an estimation tool rather than a formal inferential procedure.
Repeated K-fold Cross Validation
One potential drawback of standard k-fold cross-validation is that the results can still vary depending on how the data is split (Kuhn & Johnson, 2019). To reduce this variability, we can use repeated k-fold cross-validation. In this approach, the entire cross-validation procedure is repeated multiple times using different random partitions of the training data. This produces a larger set of performance estimates, leading to a more stable and reliable evaluation of model performance (Kuhn et al., 2022). Averaging across repeats reduces sensitivity to any single data split and improves the robustness of both the expected performance estimate and its variability.
The main drawbacks of this method are that it is computationally more expensive than standard k-fold cross-validation, since the full resampling procedure is repeated multiple times. Additionally, it does not resolve the underlying dependence between resamples. The assessment sets are still derived from overlapping training data, so the assumption of independent sampling remains violated (Lantz, 2023).
Leave-One-Out Cross-Validation (LOOCV)
A special case of k-fold cross-validation is leave-one-out cross-validation (LOOCV). In this approach, the number of folds is equal to the number of observations in the dataset. This means that in each iteration, we use all observations except one for training, and we evaluate the model on the single remaining observation.
This process is repeated for every observation in the dataset. As a result, we obtain as many performance values as there are observations. These values can again be summarized using the mean and standard deviation to estimate model performance.
LOOCV can be seen as the most extreme form of cross-validation, since it maximizes the amount of training data used in each iteration. In each step, the model is trained on almost the entire dataset, which can lead to very low bias in the performance estimate. However, this comes at a cost. Because each validation set contains only one observation, the resulting performance estimates can have high variance, especially if the dataset contains noise or outliers.
From a practical perspective, LOOCV is computationally expensive, since the model must be fitted as many times as there are observations. For large datasets, this quickly becomes infeasible, which is why LOOCV is rarely used in practice when k-fold cross-validation provides a reasonable alternative.
Monte Carlo Cross Validation
Another important resampling method is Monte Carlo cross-validation, also known as random subsampling. In this approach, we repeatedly generate random splits of the training dataset into analysis and assessment sets (for example, 80% analysis and 20% assessment). Unlike k-fold cross-validation, these splits are not mutually exclusive. This means that the same observation can appear multiple times in different assessment sets, or may not appear at all in some iterations, something that is illustrated in the plot below:
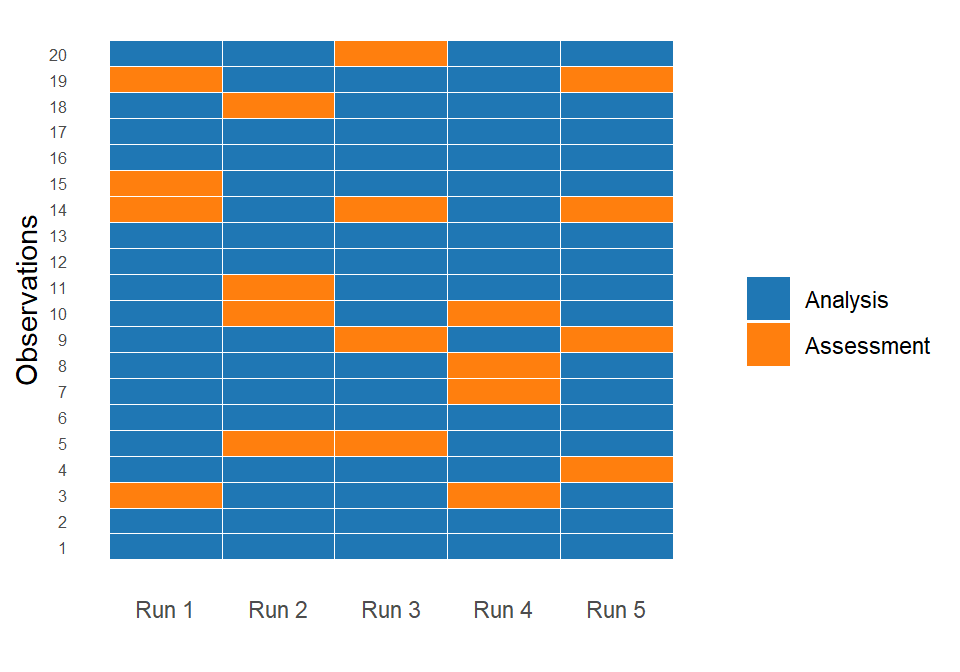
Due to overlapping splits, Monte Carlo cross-validation behaves differently from k-fold cross-validation. It provides more flexibility in how the data is split, but it does not guarantee that all observations are used equally often in validation. In return, it is useful when we want direct control over the size of the analysis and assessment sets without requiring strict partitioning structure. A typical example is evaluating a house price prediction model by repeatedly creating random 80/20 splits of the data to assess performance. In this case, we are more interested in how the model performs under different realistic samples of the data rather than ensuring that every observation is used equally often in validation, since each large split is already reasonably representative of the problem.
Due of this structural difference, Monte Carlo cross-validation and k-fold cross-validation also differ in terms of the bias–variance trade-off. In this context, bias refers to systematic error in the estimated model performance compared to the true generalization performance, while variance refers to how much the performance estimates fluctuate across different resamples.
K-fold cross-validation tends to produce lower variance estimates because every observation is used in a more structured and balanced way across folds. However, it may introduce slightly higher bias depending on the choice of K, since each model is trained on a fixed and somewhat constrained proportion of the data in every iteration. Monte Carlo cross-validation, on the other hand, can reduce bias because the random splits allow more flexibility in how training and validation sets are formed, and each split more closely resembles an independent holdout experiment. However, this comes at the cost of higher variance, since results depend more strongly on the randomness of the splits and the uneven participation of observations across iterations.
In short, Monte Carlo cross-validation tends to provide a more accurate estimate of the true generalization performance but with higher uncertainty (variance), whereas k-fold cross-validation produces more stable (lower variance) but slightly more constrained and potentially more biased estimates.
Bootstrap Resampling
In Chapter Tree-Based Models, we used bootstrap resampling for bagged trees and random forests to generate different decision trees. This same resampling technique can also be used to evaluate model performance. As discussed in that chapter, bootstrap works by repeatedly drawing samples with replacement from the original training dataset. This means that some observations may appear multiple times in a given resample, while others may not appear at all.
Each bootstrap sample is used to fit a model, and performance is evaluated on the observations that were not included in that sample. These are known as the out-of-bag (OOB) observations. By repeating this process many times, we obtain a distribution of performance estimates, similar in spirit to cross-validation.
The key difference between bootstrap and cross-validation lies in how the resamples are constructed. In k-fold and Monte Carlo cross-validation, the data is partitioned into analysis and assessment sets without replacement. In bootstrap, however, sampling is performed with replacement, which allows the same observation to be selected multiple times within a single resample.
Because of this, bootstrap samples have a different structure from the original dataset. On average, each bootstrap sample contains about 63% unique observations, while the remaining observations are left out and used for evaluation as OOB samples. This proportion arises from the sampling-with-replacement mechanism and converges to approximately 0.632 as the dataset size increases, a result that can be justified using large-sample (Law of Large Numbers) arguments. This property makes bootstrap particularly useful when the dataset is small, since it allows us to generate many alternative training sets without requiring additional data.
In terms of bias and variance, bootstrap methods often produce relatively low-bias estimates of model performance, but they can exhibit higher variance depending on the model and dataset size. This is partly because each bootstrap sample is effectively trained on approximately 63% unique observations, which introduces additional variability across resamples. However, the main strength of bootstrap lies not in point estimation alone, but in quantifying uncertainty, making it especially useful when the goal is not only to compare models but also to assess the stability of predictions.
In practice, bootstrap resampling is widely used for estimating confidence intervals, assessing model stability, and quantifying the variability of model performance. Unlike cross-validation methods, which mainly focus on estimating predictive performance, bootstrap is particularly useful when the goal is to understand how much the results would change under different samples of the data. For this reason, it is a key tool in data science, especially when data is limited or when quantifying uncertainty is an important part of the analysis.
Rolling Origin Forecasting
All the methods discussed up to now—cross-validation, Monte Carlo cross-validation, and bootstrap—assume that there is no time component in the data. In other words, they assume that observations are independent and do not follow a specific order. In practice, however, this assumption is often violated. Many datasets have a clear temporal structure, where the order of observations is essential.
A classic example is ice cream sales, where demand increases during summer and decreases during winter due to seasonality. In this case, randomly shuffling the data would break the temporal structure of the seasonal pattern and lead to unrealistic evaluation results.
For this reason, it does not make sense to fit a model on future observations and then evaluate it on past ones. In practice, models should always be trained on past data and evaluated on future data, since this reflects the real-world forecasting setting.
When there is a clear ordering in the data, it is therefore recommended to use rolling origin forecasting resampling methods (Hyndman & Athanasopoulos, 2021). In rolling origin forecasting, the idea is to respect the temporal order of the data when creating training and test sets.
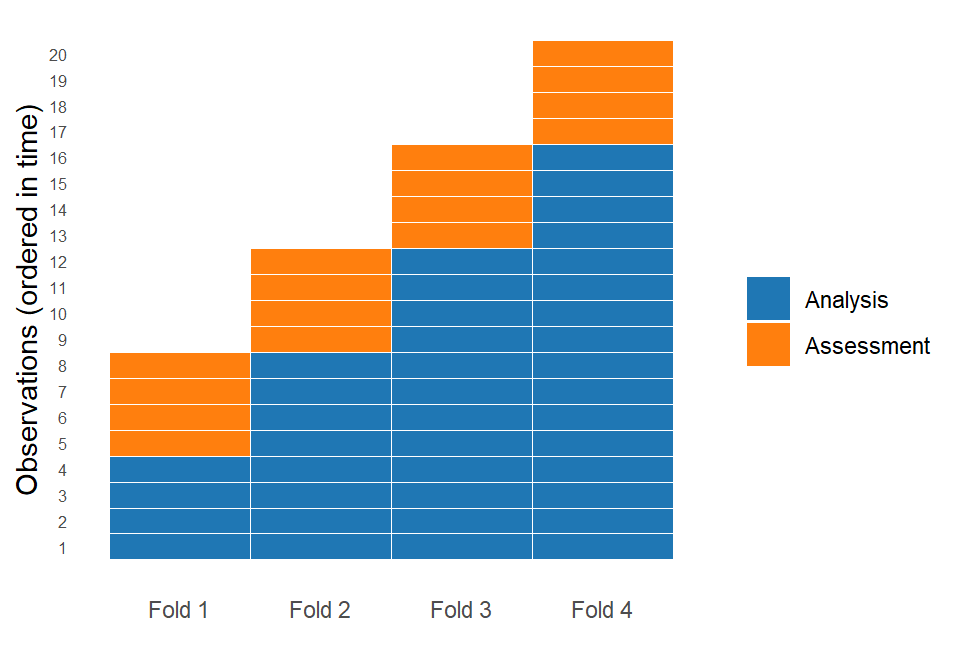
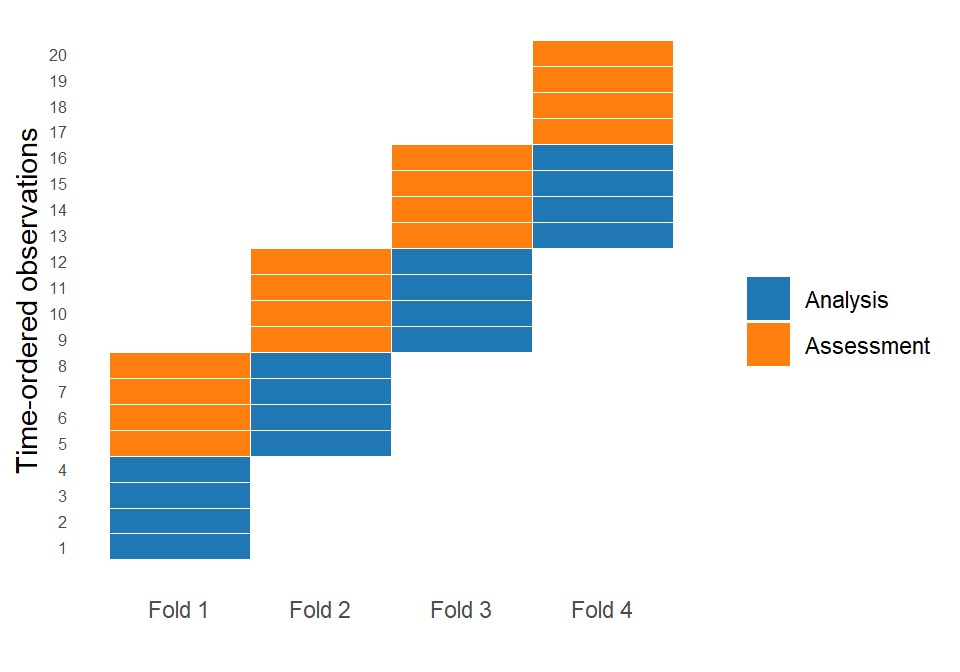
Instead of randomly splitting the data, we use either an expanding window or a sliding window approach. In an expanding window, the analysis set starts with an initial segment of the data and grows over time, meaning that all past observations are retained as new data becomes available. In a sliding window, by contrast, the analysis set has a fixed size, and as we move forward in time, older observations are removed while newer ones are added. This allows the model to focus only on the most recent data, which can be useful when older patterns become less relevant. Both approaches create splits that reflects how the model would actually be used in practice: learning from the past and predicting the future.
It is worth noting that the important element here is not strictly time itself, but the order of the observations. What matters is that the distribution of the variables may change as we move forward, a concept known as data drift. For example, customer behavior in an online store may evolve due to seasonality, marketing campaigns, or changes in preferences. Similarly, financial markets may behave differently during periods of crisis compared to stable periods.
This phenomenon is closely related to a key challenge in machine learning known as model drift. Model drift occurs when a model that performs well on recent historical data gradually becomes less accurate over time, even though it was trained on relatively recent observations. This happens because the relationship between predictors and the outcome changes, meaning that the assumptions of the model are no longer valid.
In practice, this is one of the main challenges in time-dependent data. A model that performs very well today may start to degrade tomorrow, not because it is incorrectly specified, but because the environment has changed. Rolling origin forecasting helps us evaluate models in a way that respects this evolving structure and provides a more realistic estimate of future performance.
Choosing the Appropriate Resampling Method
All the discussed resampling methods have the same goal: to produce performance estimates that generalize to previously unseen data. The next question is which method is most appropriate in each case. As always, the answer is that it depends on the problem and the data. For this reason, we need to consider how the model will be used in practice and whether there is a meaningful structure or ordering in the data.
Most of the time, our goal is to train a model using data available today to make predictions about the future. In these cases, we must respect the temporal structure of the data, and rolling origin forecasting is often the most appropriate choice, since it mimics how the model will actually be deployed in practice.
When there is no clear ordering in the data, and there is no strong evidence of model drift or distribution shift, k-fold cross-validation or repeated k-fold cross-validation are solid and widely used choices due to their balance between stability and computational efficiency, assuming a large enough dataset.
Monte Carlo cross-validation can be useful when we want more flexibility in how training and assessment sets are constructed, or when we are interested in assessing performance under many different random holdout scenarios. Bootstrap, on the other hand, is particularly appropriate when the main goal is not only prediction but also inference, such as estimating confidence intervals or quantifying uncertainty in model performance.
It is important to emphasize that all these resampling methods are applied within the training set. Best practice is to always keep a final test set separated in a “vault” that is only used once, after model selection and tuning are complete. This test set provides an unbiased estimate of final model performance. A separate test set may be omitted only in cases where the dataset is extremely small and alternative validation strategies are necessary (Kuhn et al., 2022).
Finally, once a model has been selected and validated, the most common practice is to refit it on all available training data—combining the original training, validation, and resampling information where appropriate—so that the final model benefits from the maximum amount of information before being used for future predictions (Lantz, 2023).
Performance Evaluation with Tidymodels
Now that we have discussed different resampling methods, we will use tidymodels to apply them in practice. For simplicity, we will use linear regression on the dataset eshop_revenues. Let’s start by loading the libraries and importing the dataset:
# Libraries
library(tidyverse)
library(tidymodels)
# Ensuring tidymodels functions priority
tidymodels_prefer()
# Importing eshop_revenues
eshop_revenues <- read_csv("https://raw.githubusercontent.com/Datakortex/Datasets/refs/heads/main/eshop_revenues.csv")To create an additional validation set apart from the test set, we use the function initial_validation_split(). Here, we provide the dataset and specify a two-element vector in the argument prop: the proportion for the training set and the validation set:
# Setting seed
set.seed(123)
# Splitting eshop_revenues
split_data <- initial_validation_split(eshop_revenues, prop = c(0.6, 0.2))
# Printing the results
split_data<Training/Validation/Testing/Total>
<78/26/26/130>The resulting object now contains three parts: a training set, a validation set, and a test set. To extract the corresponding tibbles, we use the functions training(), validation(), and testing() respectively:
# Extracting the training set
training_set <- split_data %>% training()
# Extracting the validation set
validation_set <- split_data %>% validation()
# Extracting the test set
test_set <- split_data %>% testing()Next, we define the linear regression model specification:
# Initiating engine
lm_spec <- linear_reg() %>%
set_engine("lm") %>%
set_mode("regression")We can now fit the model using the fit() function on the training set and evaluate performance on the validation set. We use all available predictors, with Revenue as the target variable:
# Fitting linear regression model using all predictors
lm_fit <- lm_spec %>%
fit(Revenue ~ ., data = training_set)As in the previous chapter, we generate predictions and compute the MAE. This time, however, we evaluate performance on the validation set rather than the test set:
# Making predictions on the validation set
predictions <- predict(lm_fit, validation_set)
# Calculating MAE
mean(abs(validation_set$Revenue - predictions$.pred))[1] 6603.773On average, the predicted revenues differ from the actual revenues by approximately €6,600.
We repeat the process using a more complex model that includes all available interactions between predictors:
# Fitting linear regression model using all predictors and their interactions
lm_fit_with_interactions <- lm_spec %>%
fit(Revenue ~ . * ., data = training_set)
# Making predictions on the validation set
predictions_with_interactions <- predict(lm_fit_with_interactions, validation_set)
# Calculating MAE
mean(abs(validation_set$Revenue - predictions_with_interactions$.pred))[1] 4071.508The MAE decreases substantially, suggesting that the more complex model provides better predictive performance on the validation set. Based on this result, we select this model as the final candidate, which we refit on the training and validation sets and evaluate it on the test set:
# Combining training and validation sets in a single tibble
training_validation_set <- training_set %>%
bind_rows(validation_set)
# Refitting linear regression model
lm_fit_with_interactions <- lm_spec %>%
fit(Revenue ~ . * ., data = training_validation_set)
# Making predictions on the test set
predictions_with_interactions <- predict(lm_fit_with_interactions, test_set)
# Calculating MAE
mean(abs(test_set$Revenue - predictions_with_interactions$.pred))[1] 6907.267The MAE on the test set is significantly higher than on the validation set, something that can occur when the dataset is small. This large difference illustrates an important point: performance estimates depend on the specific split of the data. For this reason, relying on a single validation split can be unstable, and motivates the use of resampling methods discussed in the previous section.
Overall, this example shows how a simple validation approach mirrors the traditional train-test procedure, while also highlighting its limitations in terms of variability and reliability.
To use resampling methods, we need to create workflows (discussed in Chapter Introduction to Tidy Modeling). Let’s keep the training set we created and define a workflow that includes the recipe of the latest model. Note that the interactions are included in the recipe using the function step_interact(). Inside the parentheses, we use a tilde (~) followed by the predictors we want to interact with each other:
# Creating recipe
lm_recipe <- recipe(Revenue ~ .,
data = training_set) %>%
step_interact(~all_numeric_predictors():all_numeric_predictors())
# Combining model and recipe into a workflow
lm_workflow <- workflow() %>%
add_model(lm_spec) %>%
add_recipe(lm_recipe)Formula Instead of Recipe
Instead of a recipe, we could actually add the formula with interactions directly in the workflow by using the function
add_formula()instead ofadd_recipe(). However, it is generally preferable to use recipes, as they offer greater flexibility and better support for structured, reproducible preprocessing workflows.
Now our workflow is ready, and we can apply resampling methods to evaluate this model. The function vfold_cv() is used for k-fold cross-validation. Even though the letter \(k\) is commonly used in the literature, tidymodels uses the letter \(v\) to denote the number of folds. We also set a seed, since cross-validation involves random partitioning of the data into folds, and we want the results to be reproducible.
The function vfold_cv() also includes the argument repeats, which is set to 1 by default. This means that standard k-fold cross-validation is performed. However, we can increase this value to perform repeated cross-validation, where the entire k-fold procedure is repeated multiple times using different random partitions of the data.
In our case, we set v = 10 and repeats = 3, which corresponds to 3 repetitions of 10-fold cross-validation:
# Setting seed
set.seed(123)
# 10x3 (repeated) cross-validation folds
folds_rep_cv <- vfold_cv(training_set, v = 10, repeats = 3)
# Printing folds
folds_rep_cv# 10-fold cross-validation repeated 3 times
# A tibble: 30 × 3
splits id id2
<list> <chr> <chr>
1 <split [70/8]> Repeat1 Fold01
2 <split [70/8]> Repeat1 Fold02
3 <split [70/8]> Repeat1 Fold03
4 <split [70/8]> Repeat1 Fold04
5 <split [70/8]> Repeat1 Fold05
6 <split [70/8]> Repeat1 Fold06
7 <split [70/8]> Repeat1 Fold07
8 <split [70/8]> Repeat1 Fold08
9 <split [71/7]> Repeat1 Fold09
10 <split [71/7]> Repeat1 Fold10
# ℹ 20 more rowsThe output is a tibble where each row represents a fold, containing an analysis set and an assessment set. In other words, each split defines which observations are used for training the model and which are used for evaluation.
Now, to fit the model across all analysis sets and evaluate it on the corresponding assessment sets, we use the function fit_resamples(). This function requires the workflow, the resampling object, and a set of performance metrics. We will discuss evaluation metrics in more detail later in this chapter, so for now we simply specify metric_set(mae) to indicate that we want to evaluate performance using MAE. In addition, fit_resamples() allows us to control several aspects of the resampling procedure via the control argument. For example, setting save_pred = TRUE inside control_resamples() stores the out-of-sample predictions for each resample, which can be useful for later inspection or diagnostic analysis:
# Fitting resamples
lm_results_rep_cv <- fit_resamples(
lm_workflow,
resamples = folds_rep_cv,
metrics = metric_set(mae),
control = control_resamples(save_pred = TRUE))
# Printing results
lm_results_rep_cv# Resampling results
# 10-fold cross-validation repeated 3 times
# A tibble: 30 × 6
splits id id2 .metrics .notes .predictions
<list> <chr> <chr> <list> <list> <list>
1 <split [70/8]> Repeat1 Fold01 <tibble> <tibble> <tibble>
2 <split [70/8]> Repeat1 Fold02 <tibble> <tibble> <tibble>
3 <split [70/8]> Repeat1 Fold03 <tibble> <tibble> <tibble>
4 <split [70/8]> Repeat1 Fold04 <tibble> <tibble> <tibble>
5 <split [70/8]> Repeat1 Fold05 <tibble> <tibble> <tibble>
6 <split [70/8]> Repeat1 Fold06 <tibble> <tibble> <tibble>
7 <split [70/8]> Repeat1 Fold07 <tibble> <tibble> <tibble>
8 <split [70/8]> Repeat1 Fold08 <tibble> <tibble> <tibble>
9 <split [71/7]> Repeat1 Fold09 <tibble> <tibble> <tibble>
10 <split [71/7]> Repeat1 Fold10 <tibble> <tibble> <tibble>
# ℹ 20 more rowsMonitoring Progress
Setting
verbose = TRUEinsidecontrol_resamples()enables progress updates during resampling, which is useful for monitoring longer computations.
The resulting object is a tibble that has list columns for the analysis and test samples, and the metric values.The resulting object is a tibble with list-columns for the analysis and test samples, as well as the corresponding metric values.
To extract the results, we use the function collect_metrics(), which returns the estimated performance metric across all folds, along with its mean and standard error:
# Collecting results
collect_metrics(lm_results_rep_cv)# A tibble: 1 × 6
.metric .estimator mean n std_err .config
<chr> <chr> <dbl> <int> <dbl> <chr>
1 mae standard 4730. 30 233. pre0_mod0_post0In our case, the output shows the average MAE across the 10 folds, as well as the variability of this estimate across different splits of the training data. This provides a more robust estimate of model performance than a single train–validation split, since it reduces the dependence on one particular partition of the data.
Since we set save_pred = TRUE earlier, we can use the function collect_predictions() to extract all individual predictions across the assessment sets. This returns a tibble containing the predictions for each resample:
# Collecting predictions
collect_predictions(lm_results_rep_cv)# A tibble: 234 × 6
.pred id id2 Revenue .row .config
<dbl> <chr> <chr> <dbl> <int> <chr>
1 24458. Repeat1 Fold01 18599. 1 pre0_mod0_post0
2 9791. Repeat1 Fold01 6241. 2 pre0_mod0_post0
3 12127. Repeat1 Fold01 13556. 20 pre0_mod0_post0
4 27139. Repeat1 Fold01 22069. 32 pre0_mod0_post0
5 31491. Repeat1 Fold01 27128. 48 pre0_mod0_post0
6 544. Repeat1 Fold01 1701. 53 pre0_mod0_post0
7 1600. Repeat1 Fold01 2927. 60 pre0_mod0_post0
8 -18.2 Repeat1 Fold01 2965. 76 pre0_mod0_post0
9 -2560. Repeat1 Fold02 367. 5 pre0_mod0_post0
10 11970. Repeat1 Fold02 18453. 17 pre0_mod0_post0
# ℹ 224 more rowsFor Monte Carlo cross-validation, we use the function mc_cv() to generate the resamples. In this case, we explicitly define two key arguments: prop, which controls the proportion of data used for the analysis (training) set in each split, and times, which determines how many random resamples are generated. Each iteration corresponds to a new random split of the data into analysis and assessment sets, independently of previous splits:
# Setting seed
set.seed(123)
# Monte Carlo cross-validation folds
folds_mc <- mc_cv(training_set, prop = 0.75, times = 30)We then use the object folds_mc instead of the k-fold cross-validation object in the fit_resamples() function. This allows us to evaluate the same model under repeated random holdout splits rather than structured folds:
# Fitting resamples
lm_results_mc <- fit_resamples(
lm_workflow,
resamples = folds_mc,
metrics = metric_set(mae))
# Collecting results
collect_metrics(lm_results_mc)# A tibble: 1 × 6
.metric .estimator mean n std_err .config
<chr> <chr> <dbl> <int> <dbl> <chr>
1 mae standard 5004. 30 187. pre0_mod0_post0Next, we move to bootstrap resampling. We use the function bootstraps() to generate bootstrap samples from the training set. In this case, the key argument is times, which controls how many bootstrap resamples are created. Each resample is generated by sampling with replacement from the original training data, meaning that some observations may appear multiple times while others may not appear at all in a given resample:
# Setting seed
set.seed(123)
# Bootstrap resamples
folds_bootstrap <- bootstraps(training_set, times = 30)We then evaluate the model using the same fit_resamples() function. In this case, performance is typically evaluated using the out-of-bag (OOB) observations, which are the data points not included in each bootstrap sample:
# Fitting resamples
lm_results_bootstrap <- fit_resamples(
lm_workflow,
resamples = folds_bootstrap,
metrics = metric_set(mae))
# Collecting results
collect_metrics(lm_results_bootstrap)# A tibble: 1 × 6
.metric .estimator mean n std_err .config
<chr> <chr> <dbl> <int> <dbl> <chr>
1 mae standard 5649. 30 239. pre0_mod0_post0These results allow us to compare different resampling strategies—k-fold cross-validation, Monte Carlo cross-validation, and bootstrap—under the same modeling framework, highlighting how different sampling methods lead to different perspectives on model performance and its variability.
Performance Metrics
Up to now, we have focused on two main metrics for measuring model performance depending on the machine learning task: accuracy for classification and mean absolute error (MAE) for regression. In practice, these metrics were mainly used for simplicity, but many alternative performance measures exist. Similar to resampling methods, the choice of the appropriate metric should depend on the goal of the analysis.
Metrics for Regression Tasks
The mean absolute error (MAE) we have used so far measures how much we expect our predictions to deviate from the actual values on average. A limitation of this metric is that it treats all errors equally, regardless of their magnitude. In many real-world applications, however, large errors are more costly than small ones. For example, in a house price prediction problem, an error of €10,000 may be acceptable when predicting a €500,000 property, but the same error would be much more problematic for a €50,000 property. In such cases, we often want a metric that penalizes large deviations more strongly. This is where the root mean squared error (RMSE) becomes useful, since it squares the errors before averaging, giving higher weight to large deviations.
\[ \text{RMSE} = \sqrt{\frac{1}{N} \sum^N_{i = 1}(y_i - \hat{y}_i) ^ 2} \]
Because of the squaring operation, RMSE penalizes large errors disproportionately more than MAE. As a result, it is more sensitive to outliers and large deviations in predictions. In practice, this also means that RMSE values are typically higher than MAE for the same model, precisely because large errors contribute much more strongly to the final score.
Another commonly used metric is the familiar R-squared (\(R ^ 2\)) from linear regression models. However, it is important to note that \(R ^ 2\) measures the proportion of variance explained by the model rather than its predictive performance on unseen data. As a result, it can be overly optimistic when evaluated on the training data or when the target variable has high variability (for example, house prices ranging from €20,000 to €5,000,000). In such cases, a high \(R ^ 2\) does not necessarily imply good out-of-sample predictive performance. For this reason, unless the goal is specifically to understand variance explained, it is generally recommended to rely on metrics such as RMSE when evaluating predictive performance (Kuhn et al., 2019).
Metrics for Hard Classification Tasks
Compared to regression tasks, hard classification tasks include a larger variety of performance metrics. As a reminder, hard classification refers to models that output a single class label rather than class probabilities. Classification problems can be either binary or multiclass. Binary classification is the most common case, and it is the focus of this section.
In binary classification, the actual outcome can only belong to one of two classes, and each prediction also assigns one of these two classes. Given the actual and predicted values, we can summarize the results in a matrix known as a confusion matrix. In this context, we assume that one class is of particular interest, which is often referred to as the event. For example, in customer churn prediction, churn would typically be considered the event of interest.
In a confusion matrix, there are four possible outcomes:
True Positive (TP): correctly predicting the event
True Negative (TN): correctly predicting the non-event
False Positive (FP): incorrectly predicting the event
False Negative (FN): incorrectly predicting the non-event
The following confusion matrix illustrates all possible outcomes:
Actual Classes | |||
Class 1 | Class 2 | ||
Predicted Classes | Class 1 | TP | FP |
Class 2 | FN | TN | |
Figure 24.5: Confusion matrix illustrating classification outcomes.
At first, the terminology can be confusing. A simple way to interpret it is that positive and negative refer to the predicted class (whether we predict the event or not), while true and false refer to whether the prediction is correct. For instance, in a medical test for a virus, a positive prediction means we predict that the virus is present. If the person actually has the virus, this is a true positive (correct prediction). If the person does not have the virus, it becomes a false positive.
Confusion Matrix Layout Conventions
The layout of the confusion matrix is not fixed. In some sources, the actual values are placed in rows and predictions in columns, while in others the opposite is used. This is purely a matter of convention; the interpretation of TP, TN, FP, and FN remains the same.
A confusion matrix allows us to compute several performance metrics. The simplest is accuracy, which we have already used in previous chapters. As discussed in Chapter K-Nearest Neighbors, accuracy is defined as the proportion of correct predictions:
\[\text{Accuracy} = \frac{\text{Number of correct predictions}}{\text{Number of observations}}\]
Using the confusion matrix notation, accuracy can also be written in the following way:
\[\text{Accuracy} = \frac{\text{TP} + \text{TN}}{\text{TP} + \text{TN} + \text{FP} + \text{FN}}\]
The interpretation is straightforward: we measure the proportion of correct predictions among all predictions. However, accuracy treats all classes equally and does not account for class imbalance. For example, if one class represents 80% of the data, a naive model that always predicts this majority class will already achieve 80% accuracy without any real predictive power.
An alternative metric is Cohen’s Kappa (Cohen, 1960; Agresti, 2012; Kuhn et al., 2019). This metric adjusts accuracy by taking into account the agreement that would be expected by chance. In other words, it compares the observed accuracy with the accuracy we would expect if predictions were made randomly according to class proportions. As a result, Kappa provides a more conservative evaluation when class imbalance is present.
Mathematically, Cohen’s Kappa is defined as:
\[\kappa = \frac{p_o - p_e}{1 - p_e}\]
where \(p_o\) is the observed accuracy (i.e., the proportion of correct predictions), and \(p_e\) is the expected accuracy under random chance agreement. The value of \(p_e\) is computed based on the marginal proportions of the confusion matrix. Intuitively, it represents how often we would expect the model and the true labels to agree purely by chance if both were generating labels independently but following the same class distribution.
To interpret Kappa, we compare the improvement over chance relative to the maximum possible improvement. A value of \(\kappa = 1\) indicates perfect agreement between predictions and actual values, while \(\kappa = 0\) indicates that the model performs no better than random guessing. Kappa can also take negative values, suggesting a performance worse than random guessing.
Intuitively, while accuracy rewards any correct prediction equally, Cohen’s Kappa asks a more strict question: “How much better is this model compared to simply guessing based on class frequencies?” This makes it especially useful when one class dominates the dataset, since a naive model can achieve high accuracy without learning meaningful patterns.
A third important metric is the Matthews Correlation Coefficient (MCC). MCC measures the correlation between the observed and predicted classifications and takes into account all four elements of the confusion matrix (TP, TN, FP, FN). Unlike accuracy and Kappa, MCC is particularly robust in the presence of class imbalance because it considers both types of errors in a balanced way. For this reason, MCC is often preferred when the dataset is highly imbalanced (Lantz, 2023).
Interestingly, if we encode the classes as 1 and 0 and compute the Pearson correlation coefficient between the true and predicted values, we obtain exactly the same result as MCC (Lantz, 2023). In this sense, MCC can be interpreted as a correlation-based measure of classification quality, capturing how strongly predictions align with the actual classes in a single value that ranges from -1 (total disagreement) to 1 (perfect agreement).
All these metrics focus on the overall performance of a classifier, combining TP and TN results in the same way. However, in many applications we are interested in distinguishing between different types of errors. Sensitivity and specificity are two metrics that focus on this distinction.
Sensitivity measures how well the model identifies the event of interest (the positive class). It is defined as:
\[\text{Sensitivity} = \frac{\text{TP}}{\text{TP} + \text{FN}}\]
Sensitivity can be interpreted as the proportion of actual positives that are correctly identified by the model. In other words, it answers the question: “Out of all real events, how many did we detect?” On the confusion matrix, this corresponds to the proportion of the positive class row that is correctly classified as TP. A perfect model would have no false negatives, meaning that all actual events are correctly detected. At the extreme, a naive model that always predicts the event would achieve perfect sensitivity, since it would never miss a positive case.
Specificity is the mirror image of specificity. It measures how well the model identifies the non-event (the negative class). It is defined as:
\[\text{Specificity} = \frac{\text{TN}}{\text{TN} + \text{FP}}\]
Specificity represents the proportion of actual negatives that are correctly classified. Intuitively, it answers: “Out of all non-events, how many did we correctly rule out?” On the confusion matrix, this corresponds to the proportion of the negative class row that is correctly classified as TN.
Together, sensitivity and specificity provide a more detailed view of model performance than accuracy alone, since they separate the ability to detect positives from the ability to correctly reject negatives.
However, in many classification problems we are primarily interested in correctly identifying the positive class (the event of interest). In these cases, we focus on metrics based on positive predictions, namely precision and recall.
Precision measures how many of the predicted positives are actually correct:
\[\text{Precision} = \frac{\text{TP}}{\text{TP} + \text{FP}}\]
It answers the question: “When the model predicts an event, how often is it correct?”. More precisely, precision focuses on the reliability of positive predictions, penalizing false alarms (FP). A high precision means that when the model predicts the event, it is usually correct.
Recall measures how many of the true positives are actually captured by the model:
\[\text{Recall} = \frac{\text{TP}}{\text{TP} + \text{FN}}\]
This formula is actually the same as sensitivity, meaning both terms are used interchangeably. The difference is mostly contextual: “recall” is more common in machine learning, while “sensitivity” is often used in statistics and medical settings.
There is typically a trade-off between precision and recall. If we make the model more “strict” about predicting positives, we reduce false positives and increase precision, but we may miss more actual positives, lowering recall. Conversely, if we make the model more “permissive” about predicting positives, we increase recall but may introduce more false positives, reducing precision.
An example that illustrates this trade-off is airplane safety classification. Suppose we have a model that predicts whether a plane is safe to depart, where a “positive” prediction represents a crash risk. On one extreme, we could predict that every plane is unsafe. In this case, recall would be perfect, because we would correctly identify all planes that would actually crash. In this case, there would be no false negatives—no planes that would actually crash would be incorrectly predicted as safe. However, precision would be extremely low, since most of our “unsafe” predictions would be false alarms—the high number of false positives would inflate the denominator in the precision formula. On the other extreme, we could predict that almost all planes are safe to fly. In this case, precision would be high, since we would only rarely predict a crash risk, and most of those predictions would be correct. However, recall would be low, as we would fail to detect the majority of actual crash cases. Note that if we predict every flight as safe (i.e., make only negative predictions), both metrics would be 0, since we would never predict a positive event, resulting in zero true positives as well as zero false positives.
This example illustrates the trade-off between precision and recall. The acceptable trade-off between these two metrics depends on the application. For instance, in medical screening we may prioritize recall, while in spam detection we may prioritize precision. A metric that combines both precision and recall is the F-measure (or F1-score). This metric uses the harmonic mean, meaning it penalizes extreme imbalance between the two metrics and only gives a high value when both are high.
The F1-score ranges between 0 and 1, where 0 indicates the worst possible performance and 1 indicates perfect precision and recall. The mathematical formula is the following:
\[\text{F1-score} = \frac{2 \times \text{precision} \times \text{recall}}{\text{precision} + \text{recall}} = \frac{2 \times \text{TP}}{2 \times \text{TP} + \text{FP} + \text{FN}}\]
In this formula, the harmonic mean ensures that the F-score will only be high if both precision and recall are high. This happens because the metric is not a simple average, but a ratio that depends on both values simultaneously.
To see this more clearly, rewrite the F1-score as:
\[\text{F1-score} = \frac{2}{\frac{1}{\text{precision}} + \frac{1}{\text{recall}}}\]
From this form, it becomes clear that if either precision or recall is small, its reciprocal becomes large, which increases the denominator and reduces the overall score. As a result, even if one metric is very high, the F-score will still be pulled down by the low value of the other.
In other words, the harmonic mean penalizes imbalance: it does not allow one metric to compensate for the other. This is why the F-score only becomes large when both precision and recall are simultaneously high.
Finally, it is important to note that the F1-score is a default choice that gives equal weight to precision and recall. In practice, this balance can be adjusted depending on the problem at hand by tuning the weight parameter (\(\beta\)). Therefore, the more general form of the F-score is given by:
\[F_\beta = \frac{(1 + \beta ^ 2) \times \text{precision} \times \text{recall}}{\beta ^ 2 \times \text{precision} + \text{recall}}\]
where \(\beta\) controls the relative importance of recall compared to precision. More specifically, values of \(\beta > 1\) place more emphasis on recall, while values of \(\beta < 1\) place more emphasis on precision. In this formula, it is apparent that when \(\beta = 1\), then \(F_\beta = F_1\).
Metrics for Soft Classification Tasks
Apart from hard classification, where predictions are assigned directly to a class, we can also use soft classification. In this case, instead of predicting a class label, the model outputs probabilities for each class. For example, rather than predicting that a customer will churn, the model may output a probability of 0.8 for churn and 0.2 for no churn. These probabilities allow us to make more flexible decisions, depending on how strict we want the classification to be.
All hard classification metrics provide a single number to evaluate performance, based on a fixed threshold (usually 0.5) that converts probabilities into class labels. However, this choice of threshold is arbitrary and may not be optimal for every application. For this reason, it is often useful to evaluate model performance across all possible thresholds, rather than relying on a single one.
The Receiver Operating Characteristic (ROC) curve is a graphical tool that illustrates the trade-off between the True Positive Rate (sensitivity) and the False Positive Rate (1 − specificity) across different classification thresholds. Instead of fixing a single cutoff value, we vary the threshold from 0 to 1 and compute the corresponding sensitivity and false positive rate at each step.
Historically, ROC curves originate from signal detection theory and were first developed during World War II to evaluate radar systems, where the goal was to distinguish between true signals (e.g., enemy aircraft) and noise. This idea was later adopted in statistics and machine learning as a way to evaluate classification models (Swets, 1988).
The following code illustrates how we can compute and visualize the ROC curve using tidymodels:
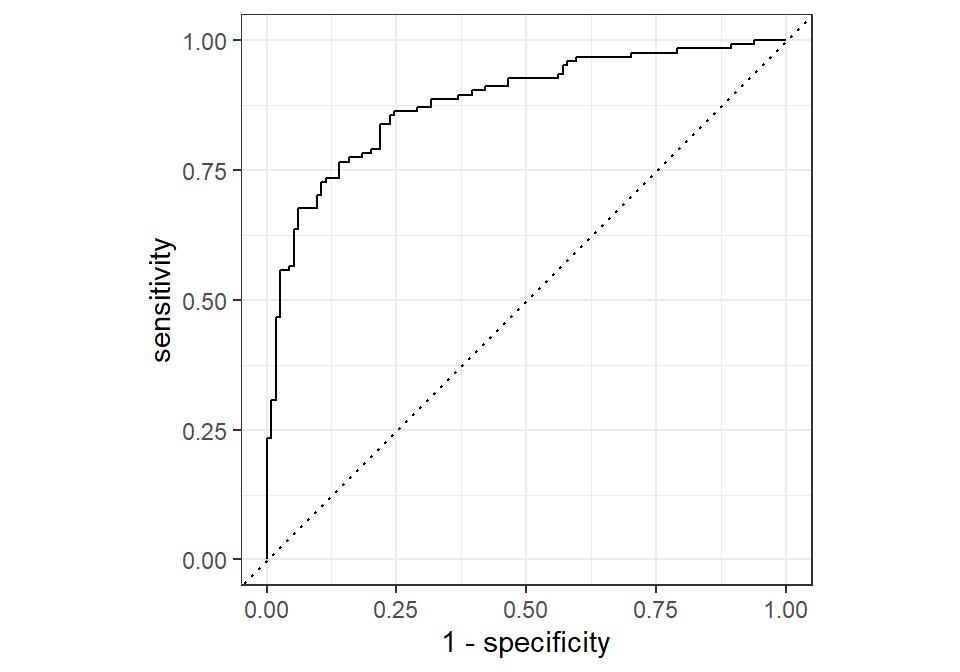
The plot shows the False Positive Rate (1 - specificity) on the x-axis and the True Positive Rate (sensitivity) on the y-axis. Each point on the curve corresponds to a different classification threshold. A model with no predictive power would produce a diagonal line, indicating that it performs no better than random guessing. In contrast, a good model will produce a curve that bends toward the top-left corner, indicating high sensitivity and low false positive rate.
In contrast to hard classification metrics, the ROC curve is less sensitive to class imbalance. This is because it focuses on rates (proportions) rather than absolute counts, making it more robust when one class is much more frequent than the other.
To summarize the ROC curve into a single number, we use the Area Under the Curve (AUC). The AUC measures the overall ability of the model to distinguish between the two classes across all possible classification thresholds. In other words, it evaluates how well the model separates positive and negative observations based on their predicted scores, rather than relying on a single cutoff value. The AUC takes values between 0 and 1:
\(0 < \text{AUC} \le 0.5\): the model performs worse than or equal to random guessing. A value close to 0.5 (e.g., 0.45) indicates random-like performance, meaning the model has no meaningful ability to distinguish between the two classes. Values below 0.5 suggest that the model is systematically ranking negative cases higher than positive ones, which can often be interpreted as an inverted model. 0.5.
\(0.5 < \text{AUC} \le 1\) : the model performs better than random guessing, with higher values indicating stronger discrimination ability. An AUC of 1 represents perfect classification, meaning that all positive observations receive higher predicted scores than all negative observations.
An intuitive interpretation of AUC is that it represents the probability that the model assigns a higher predicted probability to a randomly chosen positive observation than to a randomly chosen negative observation. Thus, higher AUC values indicate better ranking ability of the model, which is particularly useful when the goal is to prioritize or rank cases rather than make hard classification decisions. For example, if we take one patient who actually has a disease and one who does not, AUC measures how often the model assigns a higher disease probability to the sick patient than to the healthy one.
In practice, ROC-AUC is widely used because it provides a threshold-independent measure of performance. However, it should be interpreted together with other metrics, especially when the application requires prioritizing either false positives or false negatives.
In this sense, ROC-AUC can be seen as the soft classification counterpart of sensitivity and specificity, since it evaluates their trade-off across all possible thresholds rather than at a single cutoff point. Instead of fixing a threshold and computing one pair of sensitivity and specificity values, ROC analysis summarizes how these two quantities evolve together as the threshold changes.
A similar idea can be applied to precision and recall. Instead of evaluating them at a single threshold, we can construct a precision–recall (PR) curve, which illustrates the trade-off between these two metrics across different thresholds. Just like the ROC curve, each point on the PR curve corresponds to a different classification threshold.
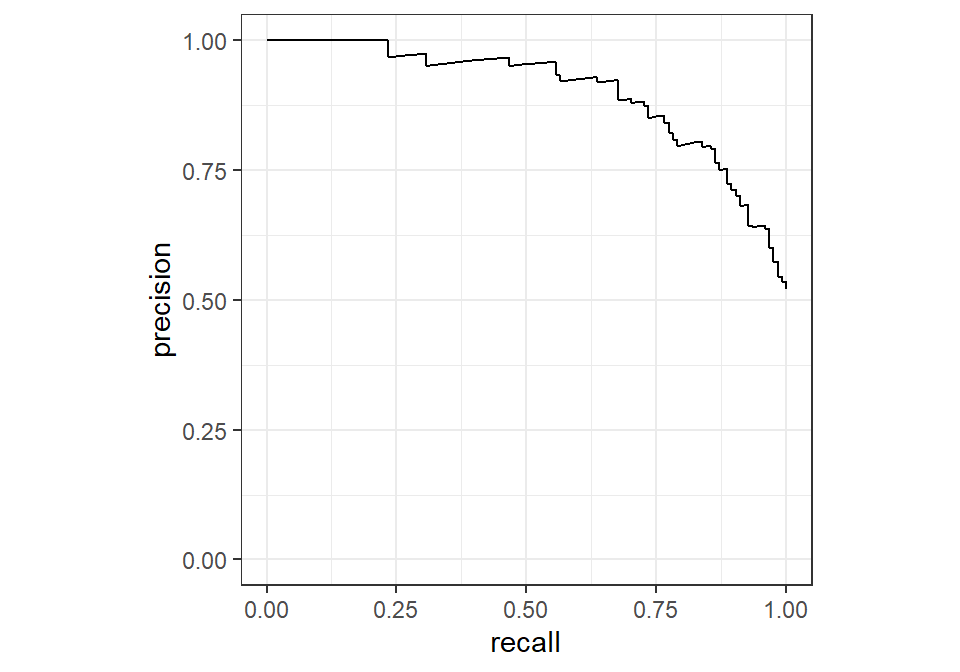
The interpretation is also similar: as we move along the curve, increasing recall typically leads to a decrease in precision, and vice versa. This reflects the same trade-off we discussed earlier, but now evaluated across all possible decision thresholds rather than at a fixed one.
In practice, the precision–recall curve is particularly useful when dealing with imbalanced datasets. While ROC curves can sometimes present an overly optimistic view in such cases, PR curves focus directly on the performance of the positive class, making them more informative when the event of interest is rare.
As with ROC-AUC, we can summarize the precision–recall curve into a single number, often referred to as the area under the precision–recall curve. This provides an overall measure of how well the model balances precision and recall across different thresholds, complementing the insights obtained from ROC-AUC.
Performance Metrics with Tidymodels
The regression and classification metrics are often harder to understand and choose appropriately than they are to implement. Especially with tidymodels, the implementation of these metrics is fairly straightforward. Tidymodels provides a consistent syntax for all metrics, something that once again illustrates its core philosophy: using the same structured approach across different modeling tasks.
The functions for the metrics discussed throughout this chapter are the following:
mae(): mean absolute errorrmse(): root mean squared errorrsq(): R-squaredconf_mat(): confusion matrixaccuracy(): overall accuracykap(): Cohen’s Kappamcc(): Matthews Correlation Coefficientsens(): sensitivityspec(): specificityprecision(): precisionrecall(): recallf_meas(): F-measure (F1-score by default)roc_auc(): ROC-AUC
All these functions require a tibble that contains the actual values and the predicted values. Typically, we specify the arguments truth for the actual values and estimate for the predicted class labels. The function roc_auc(), however, does not use the estimate argument; instead, it requires the column that contains the predicted probabilities for the positive class.
To illustrate how these metrics can be used, we load the dataset two_class_example, which is provided by the modeldata package for demonstration purposes. Since the dataset is imported as a data frame, we convert it into a tibble using the function as_tibble():
# Libraries
library(modeldata)
# Importing two_class_example
data(two_class_example)
# Transforming the imported dataset into a tibble
two_class_example <- as_tibble(two_class_example)
# Printing the first few rows
two_class_example# A tibble: 500 × 4
truth Class1 Class2 predicted
<fct> <dbl> <dbl> <fct>
1 Class2 0.00359 0.996 Class2
2 Class1 0.679 0.321 Class1
3 Class2 0.111 0.889 Class2
4 Class1 0.735 0.265 Class1
5 Class2 0.0162 0.984 Class2
6 Class1 0.999 0.000725 Class1
7 Class1 0.999 0.000799 Class1
8 Class1 0.812 0.188 Class1
9 Class2 0.457 0.543 Class2
10 Class2 0.0976 0.902 Class2
# ℹ 490 more rowsThe object two_class_example has four columns:
truth: the actual class labelsClass1: predicted probabilities for Class 1Class2: predicted probabilities for Class 2predicted: predicted class labels
The following code shows how we can create a confusion matrix and calculate all classification metrics:
# Confusion matrix
conf_mat(two_class_example, truth = truth, estimate = predicted) Truth
Prediction Class1 Class2
Class1 227 50
Class2 31 192# Accuracy
accuracy(two_class_example, truth = truth, estimate = predicted)# A tibble: 1 × 3
.metric .estimator .estimate
<chr> <chr> <dbl>
1 accuracy binary 0.838# Cohen's Kappa
kap(two_class_example, truth = truth, estimate = predicted)# A tibble: 1 × 3
.metric .estimator .estimate
<chr> <chr> <dbl>
1 kap binary 0.675# Matthews Correlation Coefficient
mcc(two_class_example, truth = truth, estimate = predicted)# A tibble: 1 × 3
.metric .estimator .estimate
<chr> <chr> <dbl>
1 mcc binary 0.677# Sensitivity
sens(two_class_example, truth = truth, estimate = predicted)# A tibble: 1 × 3
.metric .estimator .estimate
<chr> <chr> <dbl>
1 sens binary 0.880# Specificity
spec(two_class_example, truth = truth, estimate = predicted)# A tibble: 1 × 3
.metric .estimator .estimate
<chr> <chr> <dbl>
1 spec binary 0.793# Precision
precision(two_class_example, truth = truth, estimate = predicted)# A tibble: 1 × 3
.metric .estimator .estimate
<chr> <chr> <dbl>
1 precision binary 0.819# Recall
recall(two_class_example, truth = truth, estimate = predicted)# A tibble: 1 × 3
.metric .estimator .estimate
<chr> <chr> <dbl>
1 recall binary 0.880# F1-score
f_meas(two_class_example, truth = truth, estimate = predicted)# A tibble: 1 × 3
.metric .estimator .estimate
<chr> <chr> <dbl>
1 f_meas binary 0.849# ROC-AUC score
roc_auc(two_class_example, truth = truth, Class1)# A tibble: 1 × 3
.metric .estimator .estimate
<chr> <chr> <dbl>
1 roc_auc binary 0.939Regarding workflows, we can actually insert the chosen metrics in the metric argument using the metric_set() function, just like we did earlier in this chapter. To illustrate, let’s calculate MAE, RMSE and R-squared using the Monte Carlo cross-validation. The difference now is that we insert the metrics in the metric_set() function:
# Fitting resamples to calculate MAE, RMSE and R-squared
lm_results_mc <- fit_resamples(
lm_workflow,
resamples = folds_mc,
metrics = metric_set(mae, rmse, rsq))
# Collecting results
collect_metrics(lm_results_mc)# A tibble: 3 × 6
.metric .estimator mean n std_err .config
<chr> <chr> <dbl> <int> <dbl> <chr>
1 mae standard 5004. 30 187. pre0_mod0_post0
2 rmse standard 6500. 30 250. pre0_mod0_post0
3 rsq standard 0.829 30 0.0192 pre0_mod0_post0The output now consists of multiple rows, one for each metric. For each metric, we obtain the average performance across all resamples, along with measures such as the standard error. This allows us to evaluate the model from different perspectives—for example, comparing absolute errors (MAE), penalizing large errors (RMSE), and assessing explained variance (R-squared)—all within a single, consistent framework.
In practice, this flexibility is particularly useful, as different metrics may highlight different strengths and weaknesses of the same model.
Recap
In this chapter, we focused on resampling methods such as k-fold cross-validation, Monte Carlo cross-validation, and bootstrap. These methods allow us to evaluate models multiple times on different splits of the data, providing a more stable and robust estimate of performance than a single train–test split. We also highlighted how different resampling strategies reflect different assumptions about the data, including independence, randomness, and temporal structure. Additionally, we showed how these evaluation strategies connect naturally to performance metrics for both regression and classification problems.
Using tidymodels, we demonstrated how both resampling methods and performance metrics can be implemented within a unified syntax, highlighting the consistency and simplicity of the framework.